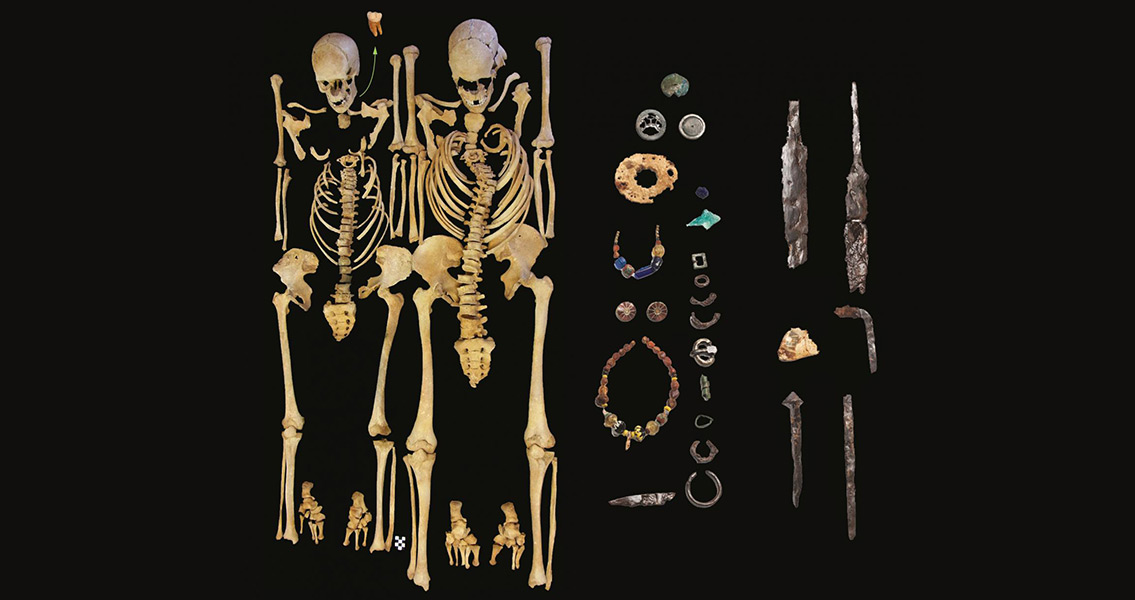
Yersinia pestis, the bacterium behind the Black Death, could also have been behind the sixth century plague epidemic. Michal Feldman, Johannes Krause and Michaela Harbeck, scientists based in Germany who are among the authors of the study, have recovered the bacterial culprit from sixth century skeletons found in Altenerding, an ancient southern German burial site near Munich. The genome found at Altenerding dates back to the beginnings of the outbreak of the plague, and confirms Yersinia pestis was the bacteria behind the outbreak. The new sequence reveals new insights into the molecular evolution of Yersinia pestis since the Byzantine period. It has also revealed new features that couldn’t be detected in a previous ‘draft genome’ of the bacterium, including 30 newly identified mutations and structural rearrangements unique to the Justinianic strain. “The fact that the archeological skeletons which gave these exciting insights were excavated over 50 years ago underscores the importance of maintaining well curated anthropological collections,” said Harbeck. “We were very fortunate to find another plague victim with very good DNA preservation in a graveyard just a few kilometers from where the individual analyzed in Wagner et al. [a recent study describing a draft genome] was found. It provided us with the great opportunity to reconstruct the first high quality genome in addition to the previously published draft genome.” Harbeck and colleagues’ results suggest that the strain of Yersinia pestis behind the Justinian Plague was more genetically diverse than previously thought. They are still unable to determine how and why the lethal pathogen reached Germany, however. Beyond simply analysing the development of a sixth century disease, the study could have far reaching applications. The authors have used the results to develop new guidelines that may improve the quality and authenticity of genetic data recovered from ancient pathogens. In addition, with plague re-emerging in certain regions of the world, the study provides a historical resource for the evolutionary changes, adaptation and human impact of the disease. “Our research confirms that the Justinianic plague reached far beyond the historically documented affected region and provides new insights into the evolutionary history of Yersinia pestis, illustrating the potential of ancient genomic reconstructions to broaden our understanding of pathogen evolution and of historical events,” said Feldman. “Our reanalysis of previous datasets stresses the importance of following strict criteria to avoid errors in the reconstruction of ancient pathogen genomes.” Image courtesy of State Collection of Anthropology and Palaeoanatomy Munich]]>